Key Takeaways:
- There are many proteins and genes that make up scientists’ list of most wanted cancer culprits. Some are more elusive than others.
- A new approach using nanoparticles may shed light on one previously undruggable protein, making it a potential target for a proven cancer drug.
- If successful, it may open the door for a new treatment for triple-negative breast cancer.
A protein that normally serves useful functions in the body like helping wounds heal and repairing damaged tissues is also high on scientists’ “most wanted” list of cancer culprits. Called STAT3, the protein has been found to be overactive in a variety of cancers — including breast cancer — driving malignant growth, survival, and metastasis. But unlike some other cancer-driving proteins, the rogue form of STAT3 has proven difficult to attack with drugs. A new study involving nanoparticles may shed light on how to make STAT3 a “druggable” target and pave the way for a new option for triple-negative breast cancer.
“STAT3 is a master regulator of gene expression, normally turning on and off within minutes, but in cancer cells it can get locked in the ‘on” position, driving high expression of genes that promote growth and invasion of tissues,” says David Frank, MD, PhD, who has been studying the protein for more than 20 years. STAT3 is overactivated in about 70 percent of breast cancers and nearly all “triple-negative” breast cancers, which are highly aggressive and for which effective treatments are lacking. Frank — and a growing number of researchers — has been seeking ways to target STAT3 in tumors.
Unfortunately, the structure of STAT3 places it among what scientists call “undruggable” proteins. For a drug to inhibit the activity of an oncogenic (cancer-causing) protein, molecules of the drug need to bind to the protein by latching onto some part of its structure. “As opposed to many other cancer proteins that have a nice ‘pocket’ for a drug to fit in, STAT3 has big flat surfaces” that don’t provide a good docking point for drugs, explains Frank.
So, Frank devised a strategy for attacking cancer cells containing overactivated STAT3 in a somewhat indirect fashion. He is the senior author of a report on the work in the journal Molecular Cancer Therapeutics.
Exploiting a difference
First, he and his colleagues discovered that the activated STAT3 protein in cancer cells altered the metabolism of lipid compounds, some of which are components of the cells’ outer membranes. As a result, the outer membranes of the STAT3-driven cancer cells were “remodeled,” or made structurally somewhat different, from the membranes of normal cells. If the scientists could find a way to exploit this difference, they might be able to target the cancer cells without having to directly confront the STAT3 protein.
Frank worked with scientists from the Massachusetts Institute of Technology who created tiny nanoparticles made up of a core, surrounded by layers of different chemicals designed to interact with the outer membranes of cells. These “layer-by-layer” nanoparticles were tested to identify which chemical coating would preferentially attach to the STAT3-altered outer membranes of breast cancer cells.
The researchers found one that bound with 50 percent greater efficiency to activated-STAT3 breast cancer cells than to normal breast cells, suggesting that the nanoparticles could be used as a safe delivery vehicle for cancer drugs.
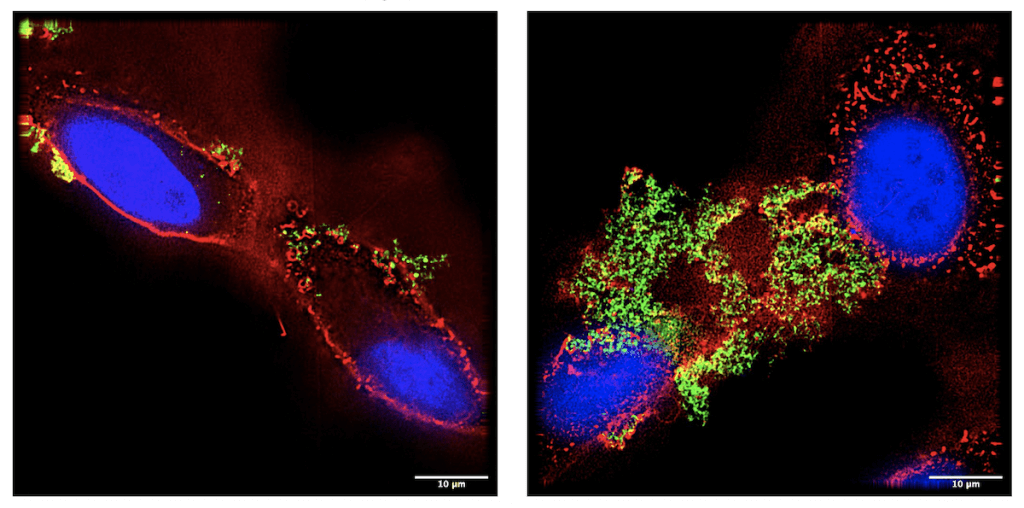
To test this strategy, the investigators loaded the nanoparticles with cisplatin — a chemotherapy drug sometimes used to treat triple-negative breast cancer. They demonstrated that cisplatin-carrying nanoparticles preferentially homed to the breast cancer cells, rather than normal cells, grown in laboratory dishes. Furthermore, the nanoparticles were able to penetrate densely packed three-dimensional breast cancer “organoids,” also in the laboratory.
The cisplatin-loaded nanoparticles were also able to kill breast cancer cells at lower doses than what is needed with traditionally delivered cisplatin. In addition, the investigators tested the effect of radiation — which is often used in treating breast cancer — and found that it increased the specific binding of the drug-carrying nanoparticles.
The next step, Frank says, is to test the system in animal models.
“These findings provide a promising starting point for the development of a rational, targeted approach to treating triple-negative breast cancers,” say the investigators.
