There are several ways to ruin a tune: play the wrong notes, play them at the wrong time, or with the wrong emphasis. Cancer cells, working from a corrupted score – their genome – manage to do all three.
In cancer, bad instructions meet improper execution. Tumor cells are notorious for alterations of their genome: mutations, misspellings of the genetic code, out-of-place stretches of DNA, missing genes, overabundant genes. But they’re also marked by changes in gene expression – the activity level of individual genes — a process controlled by the cell’s epigenetic machinery.
Instances of epigenetic changes have been found in many forms of cancer. But while scientists know a great deal about how mutations and other errors of the genome occur, the causes of epigenetic alterations in cancer remain cloudy.
In a recent study in the journal Nature, scientists at Dana-Farber describe a mechanism by which such alterations could arise — a process that involves one of the most perilous, harrowing experiences a chromosome can undergo.
‘Ruthless mutation machines’
The finding is the latest to emerge from David Pellman, MD, and his colleagues’ research into the sometimes inept way cancer cell divide, and the effect this can have on the cells’ nuclei and the chromosomes within them. They and other scientists have shown that slip-ups in cell division can lead to the formation of micronuclei — tiny compartments holding a single chromosome arm – and chromosome bridges — stretched-out chromosomes that span two daughter cells. Both micronuclei and chromosome bridges are extraordinarily fragile; when they break, it’s as though the affected chromosome is a delicate wine glass dropped on a stone floor. Its shattering, known as chromothripsis, results in extensive damage to the cell’s DNA, Pellman’s lab has shown.
The impact of this accident can reverberate across generations of cells. When a chromosome bridge collapses, its shards enter the newly formed daughter cells, where they’re sealed in micronuclei. If the micronuclei rupture, as they often do, the bits of chromosome inside may splinter further. The loss of micronuclei also triggers an inflammatory immune response. The process – and the potential for more chromosomal damage — is renewed with each round of cell division.
“It’s well established that these abnormal nuclear structures — chromosome bridges and micronuclei — act as ruthless mutation machines, impairing the chromosomes that make their way into them,” says Pellman. “For the current study, we asked whether chromothripsis also has an impact on a cancer cell’s epigenetic mechanisms — adding more insult to the injury of the genetic damage.”
There was good reason for thinking so. The battering that chromosomes undergo during chromothripsis might easily be thought to wreak havoc not only with their DNA but also with the apparatus that opens and closes the throttle on gene activity.
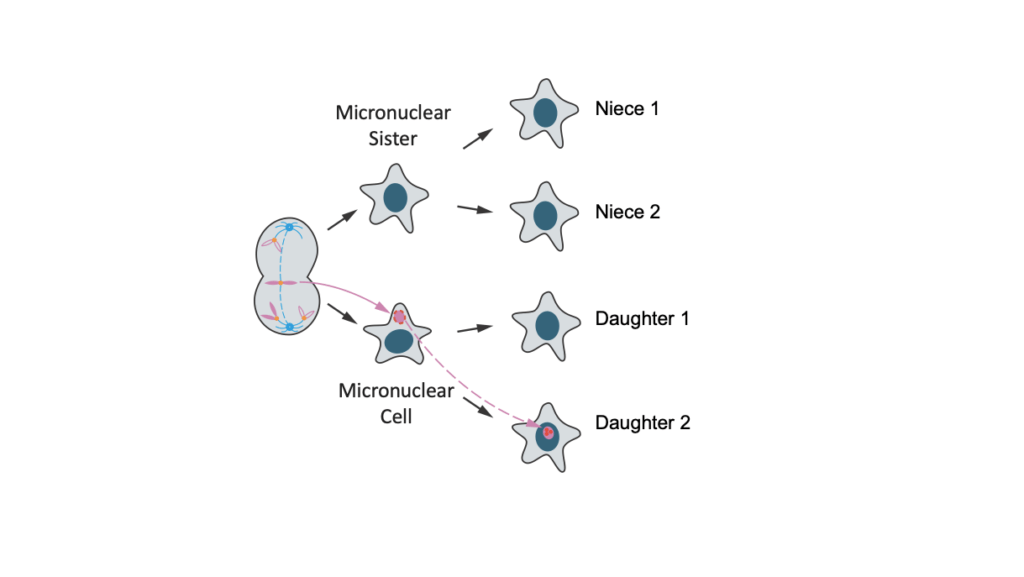
The above illustration portrays how errors in cancer cell division may cause some genes to go silent. In the left-hand image, a chromosome “lags” in the middle of the cell because of an error in cell division. The missegregated chromosome becomes trapped in a micronucleus. The membrane around the micronucleus is fragile and breaks, leading to DNA damage and silencing of gene transcription — the transfer of genetic information to RNA (middle column of cells). In the second generation, the chromosome from the micronucleus is integrated into a normal daughter nucleus (in cell at lower right) but still has defective transcription.
Addition and subtraction
Genes are rendered more or less active by a complex system that attaches and removes chemical alterations to the structure that holds DNA in place. Within the chromosome, strands of DNA are wrapped around clusters of proteins called histones like threads on a spool. The adding and subtracting of chemical subunits to the histones causes these coils to tighten or loosen. The DNA in a taut section is essentially locked away — not available for use by the cell. A looser section is like an unrolled scroll: the genetic information within it can be “read” as a template for making cell proteins.
To see whether this intricate machinery can survive chromothripsis unscathed — and, if not, whether disruptions of the machinery are passed on to later generations of cells — researchers did a series of experiments, using tools and techniques of their own creation.
Pellman’s lab, in collaboration with Dana-Farber data scientist Cheng-Zhong Zhang, PhD, began with a modified version of a technique they created nearly a decade ago to track the genetic devastation wrought by chromothripsis. The original version, called Look-Seq, enabled researchers to watch as an individual cell bungled the process of division – as a piece of chromosome slipped into the wrong daughter cell, as a micronucleus formed around it, as the micronucleus disintegrated and the bit of chromosome inside shattered. Then, by sequencing the DNA in the daughter cell, they could discover the changes that had occurred within its genome.
A catapult for RNA
For the new study, researchers created Look-Seq2, which, unlike its predecessor, sequences the RNA of the daughter cell and its progeny, rather than the DNA. RNA carries genetic information – acquired from DNA through a process known as transcription — to the protein-making factories of the cell. As a result, the set of RNA molecules within a cell offers a snapshot of which genes are active, and how active they are, at a given time.
To sample the RNA within a single cell, and the generations of cells that spring from it, the study’s first author, Stamatis Papathanasiou, PhD, of Pellman’s lab, and co-author and co-patent holder Hauibin Zhang, PhD, of the Broad Institute and Harvard designed and 3D-printed a device for use with technology known as laser capture microdissection.
Pellman explains how it works: “A group of cells are arrayed in a membrane on a glass slide. We use a laser to cut out the membrane around individual cells so they aren’t damaged. The DNA isn’t disturbed when the cells are isolated during the original Look-Seq procedure, but their transcription state — their pattern of gene expression — might change if they’re irritated too much. So, in the updated approach, we cut the cells out and use the catapulting force of the laser to propel it onto a plate with 384 microscopic wells. We use imaging technology to identify the cells with micronuclei and sequence their RNA.”
With this information in hand, researchers faced a new riddle: how to know whether the transcription patterns in cells with micronuclei were the same as, or different from, those in normal cells. To answer it, they employed a bioinformatics technique developed by Cheng-Zhong Zhang, which calculates the expected pattern of transcription in cells with intact chromosomes and compares it to that in cells with abnormal nuclear structures like micronuclei and chromosome bridges. Sequencing the micronucleated cell’s family members allows the expected pattern of transcription to be inferred. With this technique, investigators could determine not only whether there were epigenetic changes in these cells but also whether those changes persisted into subsequent generations of cells.
A radical alteration
The researchers found that in micronucleated cells, gene activity within chromosomes surrounded by micronuclei had essentially ceased. When researchers analyzed the cells’ epigenetic marks – the arrangement of chemical subunits on histones — they found these to be radically altered from those on normal cells.
“Our analysis of the marks was consistent with the idea that the chromosomes are largely inactivated by the trauma of being exposed to the cytoplasm of the cell when the micronucleus ruptures,” Pellman states.
When cells with micronuclei — or that once had micronuclei — divide, the chromosomes from those micronuclei can be incorporated into the nucleus of one of the daughter cells, where a molecular spell-check system corrects snafus in their DNA. Pellman and his colleagues wanted to know whether epigenetic snafus are corrected as well, or whether they persist in subsequent generations of cells.
The answer, it turns out, is a little of both. Whereas nearly all the genes in the chromosome from the parent cell may be silenced, roughly 30% of them are silenced in a daughter cell. “We showed that while epigenetic changes can be inherited by a daughter cell, it’s not an all or nothing phenomenon,” Pellman remarks. “The extent of that inheritance is variable.”
As a result, each generation of cells may inherit a different set of epigenetic alterations. In tumor cells, some of these changes will be helpful, supporting their growth and proliferation, and others won’t. Cells that inherit the most advantageous changes will be the most likely to survive.
“Our findings show that the well-known instability of the chromosomes in cancers is inherently coupled to a degree of instability of the epigenome,” says Pellman. “This identifies yet another shape-shifting feature of cancer cells — helping explain their ability to rapidly evolve and evade drug treatments and the immune system.” A study by Samuel Bakhoum, MD, PhD, and colleagues from the Memorial Sloan Kettering Cancer Center reached similar conclusions using different approaches and is published together with the paper from Pellman and Zhang.
